Research project
T-DNA integration and DNA repair of DSBs in plants
Identification and characterization of components of DNA repair pathways and their role in Agrobacterium T-DNA integration and repair of CRISPR/Cas induced DSBs.
- Duration
- 2011 - 2017
- Contact
- Sylvia de Pater
- Funding
-
 Chinese Scolarship Counsel
Chinese Scolarship Counsel
-
 KNAW
KNAW

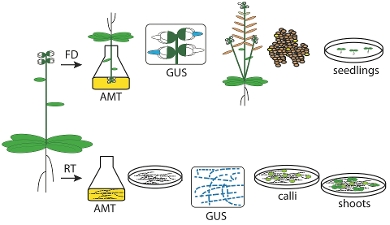
In eukaryotic organisms DNA double strand breaks (DSB) are repaired via two mechanisms: homologous recombination (HR) and non-homologous end-joining (NHEJ). Since there are several different NHEJ pathways, we want to identify components of these repair pathways in order to inactivate them or manipulate the balance between HR and NHEJ in favor of gene-targeting via HR. Arabidopsis DNA repair mutants are being used and by crossing several multiple mutations in different repair pathways are combined. T-DNA integration is being analysed in these DNA repair mutants. One essential protein for T-DNA integration is Polθ (van Kregten et al 2016). This protein was first identified in animal models, but is also conserved in plants, where is creates characteristics scars at the T-DNA – plant DNA junction. Repair of CRISPR/Cas-induced DSBs in the various mutants showed that longer deletion and more severe mutations are obtained when Ku80, a subunit of a complex that protects the ends of DSBs, is mutated.
- Jia Q, Bundock P, Hooykaas PJJ and de Pater S (2012) Agrobacterium tumefaciens T-DNA integration and gene-targeting in Arabidopsis thaliana non-homologous end-joining mutants. J Bot, Article ID 989272, 13 pages
- Jia Q, den Dulk-Ras A, Shen H, Hooykaas PJJ and de Pater S (2013) Poly(ADP-ribose)polymerases are involved in micro-homology mediated back-up non-homologous end-joining in Arabidopsis thaliana. Plant Mol Biol 82, 339-351
- Van Kregten M, de Pater s, Romeijn R, van Schendel R, Hooykaas PJJ and Tijsterman M (2016) T-DNA intergration in plants results from polymerase-θ-mediated DNA repair. Nature Plants doi: 10.1038/NPLANTS.2016.164
- Shen H, Strunks GD, Klemann BJPM, Hooykaas PJJ, de Pater BS (2017) CRISPR/Cas9-induced double-strand break repair in Arabidopsis nonhomologoud end-joining mutants. doi: 10.1534/g3.116.035204
