Research project
Personalized Medicine
Getting personal
- Contact
- Laura Heitman
Genetic variation between individuals can have a large effect on drug action and effectiveness. Each year, over a hundred billion dollars and Euros are spent on drugs that don’t help the patients taking them. Even the most popular 'blockbuster' drugs work in merely 35-75% of patients. Patients differ in response to drugs through differences in genes, environment and lifestyle. Personalized medicine or precision medicine aims to use the patient’s genetic information to fine-tune drug prescriptions, thereby decreasing the risk of ineffective treatment, inappropriate dosing or side-effects. New technologies, such as label-free cell-based assays, are creating opportunities to make the search for medicine personalization cheaper and easier.
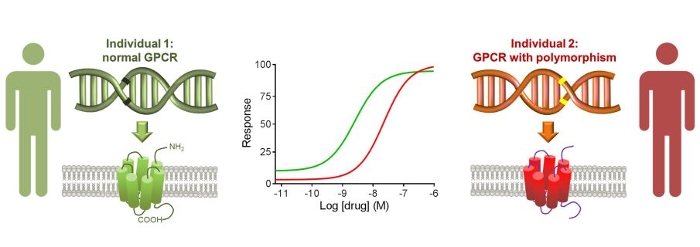
In order to determine whether different genetic variants of drug targets indeed leads to differences in drug responses, our collaborative project between the VU, LACDR and LUMC uses a combination of bioinformatic tools and novel label-free, cell-based high throughput screening methods. The project focuses on G protein-coupled receptors (GPCRs), as these form the target for almost 50% of current marketed drugs. To assess the viability of the personalized medicine approach for GPCRs, the two main questions that we want to answer are: 1) whether different drug responses are observed when different variations are detected in the corresponding drug target pathway; 2) whether similar drug responses are observed when identical variations are detected in the corresponding drug target pathway.
To investigate how genetic variation in GPCRs lead to differences in drug response, personal cell lines could offer an ideal model system. We have developed a methodology to characterize GPCR responses in a type of personal cell line using a novel, label-free cellular assay. We optimized and applied the methodology to study the genetic effects on the function of several GPCRs already, for instance the cannabinoid receptor 2 and adenosine receptor A2A. In both cases we were able to identify compounds that showed genotype-related effects, such as a partial agonist on the A2AR, as well as identify ‘blockbuster’ drug candidates that are less prone to individual differences. Hence, this methodology opens up the possibility to use such personal cell lines as a model system for a very direct investigation of the influence of genetic variation on drug response.
Hillger JM, Schoop J, Boomsma DI, Eline Slagboom P, AP IJ, Heitman LH. 2015. Whole-cell biosensor for label-free detection of GPCR-mediated drug responses in personal cell lines. Biosens Bioelectron 74:233-242.
Hillger JM, Diehl C, van Spronsen E, Boomsma DI, Slagboom PE, Heitman LH, AP IJ. 2016. Getting personal: Endogenous adenosine receptor signaling in Lymphoblastoid Cell Lines. Biochem Pharmacol doi:10.1016/j.bcp.2016.06.006.
