Research project
Innate immune defence against intracellular pathogens
What are the host immune defence mechanisms that control intracellular infections and how are these subverted by pathogens?
- Duration
- 2004 - 2016
- Contact
- Annemarie Meijer
- Funding
-
 European Commission 7th framework programme Grant Agreement ZF-Health
European Commission 7th framework programme Grant Agreement ZF-Health
-
 European Commission 7th framework programme Grant Agreement FishForPharma
European Commission 7th framework programme Grant Agreement FishForPharma
-
 Smart Mix Program of the Netherlands Ministry of Economic affairs and the Ministry of Education
Smart Mix Program of the Netherlands Ministry of Economic affairs and the Ministry of Education
-
 ZonMW EnablingTechnologies
ZonMW EnablingTechnologies
-
 CSC Chinese government scholarships
CSC Chinese government scholarships
-
 HEC Higher education commission of Pakistan
HEC Higher education commission of Pakistan
-
 European Commission Horizon 2020 programme
European Commission Horizon 2020 programme
- Partners
Prof. dr. Steven Renshaw, University of Sheffield
Dr. Philip Elks, University of Sheffield
Dr. Annette Vergunst, Université de Montpellier
Dr. Gillian Tomlinson, University College London
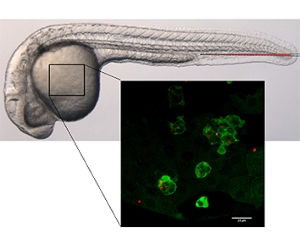
We use zebrafish models to study the role of evolutionary conserved processes involved in immunity and inflammation during bacterial and fungal infections.
We use zebrafish models for studying innate host defence mechanisms involved in several bacterial and fungal infectious diseases, including tuberculosis, salmonellosis, staphylococcal infections, and aspergillosis. The major cell types of the innate immune system, macrophages and neutrophils, develop in zebrafish embryos as early as one day post fertilization, whereas development and maturation of the adaptive immune system is a process of several weeks. For this reason and because of a number of practical advantages for microscopic imaging and genetic approaches, the zebrafish embryo provides an excellent model to study specific interactions between pathogens and innate immune cells in vivo. For many years we are studying the host defence pathways activated by pattern recognition receptors, in particular Toll-like receptor (TLR) signaling. We have made extensive use of RNA sequencing to increase insight into the host immune response genes that are activated at specific stages of infection and have developed zebrafish models for immunodeficiency (mutation of the TLR adaptor Myd88) mutants) and hyperinflammation (knockdown of the regulatory phosphatase Ptpn6/Shp1). We identified the autophagy modulator Dram1 as a new connection between pathogen recognition and autophagic host defense and are studying several other innate immune genes with functions in immune cell migration (chemokine receptors of the Cxcr3 family), phagocytosis (scavenger receptor Marco), and anti-bacterial mechanisms (perforin-like proteins of the Mpeg1 family). The ultimate aim of these studies is to identify potential targets for novel host-directed therapies against infectious diseases.
- Benard EL, Racz PI, Rougeot J, Nezhinsky AE, Verbeek FJ, Spaink HP, Meijer AH (2015) The macrophage-expressed perforins Mpeg1 and Mpeg1.2 have anti-bacterial function in zebrafish. J Innate Immun 7:136-52
- van der Vaart M, Korbee CJ, Lamers GEM, Tengeler AC, Hosseini R, Haks MC, Ottenhoff THM, Spaink HP, Meijer AH (2014) The DNA Damage-Regulated Autophagy Modulator DRAM1 links mycobacterial recognition via TLR-MYD88 to autophagic defense. Cell Host Microbe 15:753-67
- van der Vaart M, van Soest JJ, Spaink HP, Meijer AH (2013) Functional analysis of a zebrafish myd88 mutant identifies key transcriptional components of the innate immune system. Dis Model Mech 6:841-54
- Kanwal Z, Zakrzewska A, den Hertog J, Spaink HP, Schaaf MJ, Meijer AH (2013) Deficiency in hematopoietic phosphatase Ptpn6/Shp1 hyperactivates the innate immune system and impairs control of bacterial infections in zebrafish embryos. J. Immunol. 190: 1631-1645
