Research project
Galactofuranose biosynthesis in the filamentous fungus Aspergillus niger
This project aims to further understand the molecular details related to the biosynthesis and function of Galf containing glycoconjugates in fungal Aspergillus spp.
- Duration
- 2010 - 2015
- Contact
- Arthur Ram
- Funding
-
 NWO - STW funded project
NWO - STW funded project
- Partners
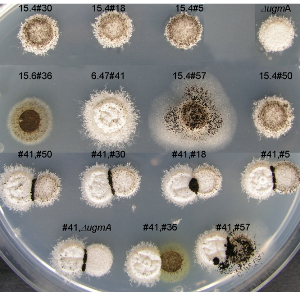
This project aims to further understand the molecular details related to the biosynthesis and function of Galf containing glycoconjugates in fungal Aspergillus spp. Galf deficient mutants are generally hypersensitive to drugs, exhibit a constitutive osmotic stress phenotype, and /or have an attenuated virulence. Since Galf has never been found in mammals and higher plants, Galf-biosynthetic pathways have raised much interest as targets for drug development to combat microbial infections.
The cell wall of filamentous fungi is an essential part of the cell protecting its integrity, forming a barrier with the surrounding environment and permitting interactions with possible substrates and hosts to be infected. Recent research has revealed that galactofuranose is essential for the integrity of the cell wall and is required for proper morphology. In its absence the mycelium grows much slower and forms abnormal hyphae with increased branching.
Galactomannans as well as O- and/or N-linked glycans on cell wall proteins and secreted (heterologous) proteins contain galactofuranose, thus showing that galactofuranose is present in different types of fungal glycoconjugates. Up to now, detailed knowledge is lacking about the galactofuranose metabolism and the role that the individual galactofuranose-containing glycoconjugates play in the integrity of the fungal cell wall. It is also unknown whether galactofuranose-containing glycoconjugates contribute to host defense mechanisms against pathogenic fungi. Research on the pathogenicity of e.g. Aspergillus fumigatus has shown that galactomannans are highly antigenic and they are therefore used to diagnose Aspergillosis in patients.
This proposal describes new strategies to identify and characterize the enzymes involved in galactofuranose metabolism. The identified genes will be used to construct mutants defective for galactofuranose biosynthesis, which will allow us to determine the function of different galactofuranose containing cell-components for cell wall integrity. Deciphering molecular details related to galactofuranose biosynthesis in Aspergillus niger to provide new opportunities for industrial applications in the field of red and white biotechnology.
- Tefsen B, Ram AF, van Die I, Routier FH. (2012). Galactofuranose in eukaryotes: aspects of biosynthesis and functional impact. Glycobiology 22 :456-469.
- Tefsen B, Lagendijk EL, Park J, Akeroyd M, Schachtschabel D, Winkler R, van Die I, Ram AF. (2012) Fungal α-arabinofuranosidases of glycosyl hydrolase families 51 and 54 show a dual arabinofuranosyl- and galactofuranosyl-hydrolyzing activity. Biol Chem. 393(8):767-775.
- Park JP, Tefsen B, Arentshorst M, Lagendijk E, van den Hondel CAMJJ, van Die I, Ram AF. Identification of the UDP-glucose-4-epimerase required for galactofuranose biosynthesis and galactose metabolism in A. niger. Fungal Biology and Biotechnology 2014, 1:6.
- Chiodo F, Marradi M, Park J, Ram AF, Penadés S, van Die I, Tefsen B. 2014. Galactofuranose-coated gold nanoparticles elicit a pro-inflammatory response in human monocyte-derived dendritic cells and are recognized by DC-SIGN. ACS Chem Biol. 9(2):383-9.
- Park J, Tefsen B, Heemskerk MJ, Lagendijk EL, van den Hondel CA, van Die I, Ram AF. 2015. Identification and functional analysis of two Golgi-localized UDP-galactofuranose transporters with overlapping functions in Aspergillus niger. BMC Microbiol. 15(1):253. doi: 10.1186/s12866-015-0541-2.
- Park J, Hulsman M, Arentshorst M, Breeman M, Alazi E, Lagendijk EL, Rocha MC, Malavazi I, Nitsche BM, van den Hondel CA, Meyer V, Ram AF. Transcriptomic and molecular genetic analysis of the cell wall salvage response of Aspergillus niger to the absence of galactofuranose synthesis. Cell Microbiol. 2016 Sep;18(9):1268-84. doi: 10.1111/cmi.12624.
