Research project
Efficient targeting of the Trichoderma genome for industrial protein engineering
The research is aimed at development of an efficient gene targeting method that allows controlled integration of DNA at a preselected site in the Trichoderma genome.
- Duration
- 2013 - 2014
- Contact
- Arthur Ram
- Funding
-
 Kluyver Centre for Genomics of Industrial Fermentation
Kluyver Centre for Genomics of Industrial Fermentation
- Partners
Dr. Igor Nikolaev and Dr. Sharief Barends - Dupont International
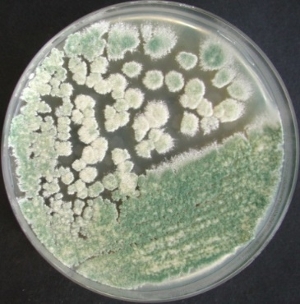
Trichoderma reesei can secrete large amounts of extracellular protein (up to 100 g/L), which makes it a paradigm host for homologous and heterologous protein production. To improve production, activity or other properties of industrially interesting enzymes, both random and systematic approaches to generate enzyme variants are employed.
To allow comparison among enzyme variants or different homologues expressed in filamentous fungi and, in particular T. reesei, it is highly desirable that the corresponding DNA constructs are integrated at a defined locus in the genome. Random integration of expression cassettes often leads to significant variation in production levels caused by differences in copy number and/or sites of integration. Therefore, this proposal is aimed at increasing the efficiency of gene targeting in T. reesei by making use of the SceI meganuclease enzyme of Saccharomyces cerevisiae to generate a DNA double-strand break (DSB) at a specific locus of the genome.
We will introduce I-SceI recognition sites into the genome of T. reesei and will investigate the effect of DSBs mediated by heterologous expression of I-SceI on transformation efficiency, on frequency of targeted integration of a reporter gene and on the expression levels of the reporter protein.
- Ouedraogo JP, Arentshorst M, Nikolaev I, Barends S, Ram AF. 2015. I-SceI-mediated double-strand DNA breaks stimulate efficient gene targeting in the industrial fungus Trichoderma reesei. Appl Microbiol Biotechnol. 99(23):10083-10095.
- Ouedraogo JP, Arentshorst M, Nikolaev I, Barends S, Ram AF. I-SceI enzyme mediated integration (SEMI) for fast and efficient gene targeting in Trichoderma reesei. J Biotechnol. 2016 Mar 20;222:25-8. doi: 10.1016/j.jbiotec.2016.02.012.
