Research project
New Methods for (f)MRI Analysis
Analysis of neuroimaging data requires multiple steps where statistics play a crucial role. The MRI methods research group develops new statistical methods that are accurate, transparent and easy to use.
- Contact
- Wouter Weeda
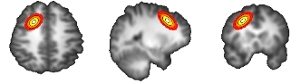
Magnetic Resonance Imaging (MRI) is a widely used method for exploring the workings of the human brain. The analysis pipeline of an MRI experiment contains multiple steps where statistics play a crucial role, often requiring researchers to make choices regarding the settings to be used. These choices can have profound impact on the outcomes of the analysis. The analysis is further complicated by the huge amount of data gathered in MRI experiments, making interpretation and quality control often difficult.
The main focus of the MRI methodology and statistics group is to clarify and improve current analysis methods and to develop new analysis methods that are both insightful and easy to use. We have four main research lines:
- Using spatial models to improve estimation of fMRI activity (Wouter Weeda)
- Improving estimation of trial-to-trial fMRI activation (Wouter Weeda)
- Improving data-driven analysis methods to identify resting-state networks (Tom Wilderjans)
- Combining multiple modalities to improve clinical prediction accuracy (Serge Rombout)
Wouter Weeda: Using spatial models to improve estimation of fMRI activity
FMRI analysis is usually performed in a massively univariate way. The temporal model is applied to every voxel in the analysis separately, requiring a post-hoc correction for multiple comparisons. Additionally, this way the spatial structure of fMRI data (that it tends to cluster in ‘blobs’ of activation) is not incorporated in the analysis. We have developed new methods that explicitly use spatial information to improve detection and interpretation of fMRI activity. These methods assume that the spatial structure of fMRI activity can be ‘captured’ by a spatial model covering a set of voxels. This creates a more sparse representation of activity, leading to improved power to detect activation and more intuitive interpretation of the spatial structure of activity. Currently we are improving these methods in terms of estimation speed and interpretability and we are designing new methods to improve connectivity analyses and multi-voxel pattern analysis. New methods are available as free R-packages.
References:
- Weeda W.D., Huizinga H.M., Christoffels I.K. & Waldorp L.J. (2009) Activated region fitting: A robust high power method for fMRI analysis using parameterized regions of activation. Human Brain Mapping 30(8): 2592-2605
List of available software:
Weeda W.D., De Vos F., Waldorp L.J., Grasman R.P.P.P. & Huizenga H.M. (2011), arf3DS4: An integrated framework for localization and connectivity analysis of fMRI data, Journal of Statistical Software 44(14): 1-33.
Wouter Weeda: Improving estimation of trial-to-trial fMRI activation
FMRI research is shifting from a functional specialization approach (where every brain area performs a specific function) to a functional integration approach (where a network of brain areas defines functionality). Given the complexity of fMRI activity - especially in event-related designs where stimulus presentations are spaced closely in time - the analysis of the brain’s response over time is often difficult. We are developing methods to improve estimation of (functional) connectivity (estimation of the network of brain areas and their interrelation) by incorporating spatial information and using more robust methods to track fMRI activity over time.
References:
- Weeda W.D., Waldorp L.J., Grasman R.P.P.P., Van Gaal S. & Huizenga H.M. (2011). Functional connectivity analysis of fMRI data using parameterized regions-of-interest, NeuroImage 54(1): 410-416.
Tom Wilderjans: Improving data-driven analysis methods to identify functional connectivity patters (networks) in resting-state fMRI data
Fundamental for brain functioning are dynamic interactions between brain regions, which are organized at large scale in functional networks and can be identified by performing Independent Component Analysis (ICA) on resting-state fMRI data. Abnormalities in functional networks have been demonstrated to be related to various neurocognitive and psychiatric disorders (e.g., dementia, panic disorder, major depression and social anxiety disorder). Data-analysis techniques to identify networks, however, often do not take the large heterogeneity in fMRI data between patients into account, which may obscure important disorder related functional network.
In our group, we are working on methods to detect functional networks that are able to also model patient heterogeneity. To this end, we combine unsupervised methods, like clustering, with existing methods for identifying networks, like ICA. As such, we hope to detect disorder related functional networks in a data-driven way.
Although we focus on fMRI data, we also plan to include other MRI modalities (i.e., multimodal integration), like information on brain structure and anatomical connections.
Serge Rombouts: Combining multiple modalities to improve clinical prediction accuracy
Early diagnosis of brain-related diseases like dementia is of vital importance in prevention and treatment. Accurately predicting at-risk groups that might develop a disease requires very sensitive and specific methods. This research line develops methods combining multiple modalities (e.g., structural, functional, resting-state) to accurately predict these at-risk groups.
Link:
