Research project
PRIORITY: Prioritising novel genomic and chemical space via federated learning and preclinical development
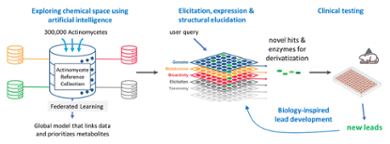
There are millions of gene clusters that encode the biosynthesis of bioactive natural products, with the vast majority yet unknown. How can we prioritize them to discover new chemical space for antibiotics? For this we will set up a pipeline based on federated learning (AI), synthetic biology and bioactivity screening and make use of a collection of >300.000 actinomycetes.
- Duration
- 2025 - 2030
- Contact
- Gilles van Wezel
- Funding
-
 NWO
NWO
- Partners
- Wageningen University
- LUMC
- NAICONS
- Fundacion Medina
- GARDP
- KU Leuven & Vlaams Instituut voor Biotechnology,
- The Wertheim UF Scripps Institute
- US Department of Energy

Description
Antibiotic resistance is a major societal problem, and the rise of multi-resistant bacteria is often referred to as the silent pandemic. However, there likely is still a vast amount of 'chemical space' yet to be discovered. Actinomycetes produce two-thirds of all antibiotics, and the PRIORITY project aims to discover new (families of) molecules and improve existing drug candidates using artificial intelligence and advanced analytical methods, as the basis for our future antibiotics.
We aim to make these medicines available to all countries, regardless of their economic prosperity. We do so with a strong global network of public and private partners.