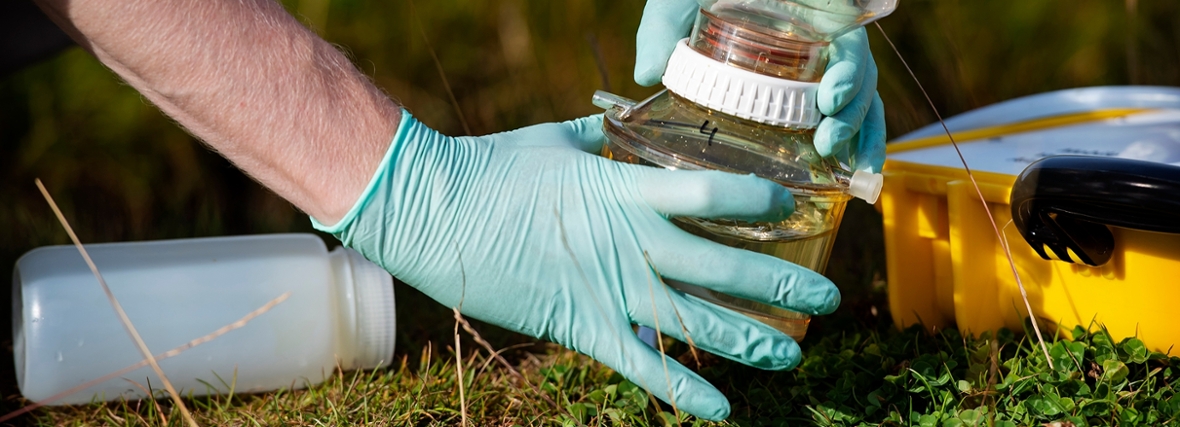

Which DNA is floating in the ditch?
You pour a scoop of ditch water in the DNA scanner, and voilà: you know exactly which plants and animals the ditch accommodates. Well, it is not that simple yet, but according to PhD candidate Kevin Beentjes, we can already use DNA techniques to monitor the quality of freshwater. For his PhD research, he investigated how we can do this faster and cheaper. Promotion on 8 April.
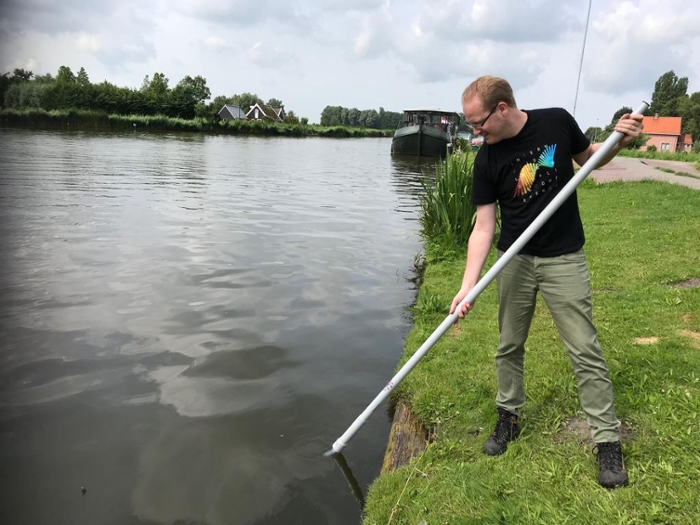
With big jackboots and a fishing net between the water plants: that is how biologists currently determine which animal species live in a ditch or pond. ‘The presence or absence of certain species says something about the water quality,’ says Beentjes. ‘At present, we still determine this mainly by hand. We count large animals on the spot, the smaller ones we collect to identify one by one under the microscope.’
The work is quite labour-intensive, and a biologist collecting animals also upsets the life in the ditch. ‘Besides, the outcome also depends on the biologist who assesses the sample’, adds Beentjes. ‘A beetle expert does not recognize worms very well and vice versa.’ The use of DNA technology could change all that.

DNA barcodes and sample smoothies to identify animals
Beentjes explains that DNA can be used in two ways to monitor water quality. 'In both cases, we use so-called DNA barcodes. These are short pieces of DNA that are specific to a certain species. Using an enormous database, we can look up which animal belongs to which barcode.'
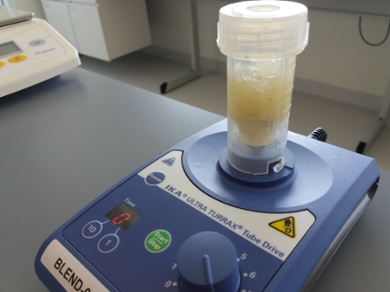
First, the DNA barcodes can be used to identify the manually collected animals. 'We put all the samples in a blender,' says Beentje. 'From the resulting smoothie we extract the DNA and compare it with the references in the database.' This method is particularly useful for animal species that are difficult to distinguish from each other at first sight, such as different species of chironomids or tiny plankton. However, for some species, such as snails, the DNA is not yet distinct enough, and other groups are still poorly represented in the database. For these groups, DNA is not yet widely applicable.
'We found about as many species as in the traditional way, but the results did not overlap completely. So we found partially different species with the different methods. I therefore think that DNA identification cannot yet directly replace the traditional method, but the methods could complement each other nicely.'

DNA fragments in the water already give a good indication
It would be even better if you could skip the collection step altogether. Beentjes: 'Every organism leaves behind traces of DNA, for example by shedding its skin, through excrement or during reproduction. We call this environmental DNA or eDNA. By taking a water sample and filtering out the eDNA, biologists can use the DNA barcodes to determine which animal species live in that water. Again, for some species this already works quite well. Especially for the larger species that leave behind a lot of DNA fragments, according to Beentjes. 'But a beetle, covered with hard armour, hardly loses any body material and is therefore much more difficult to find.'
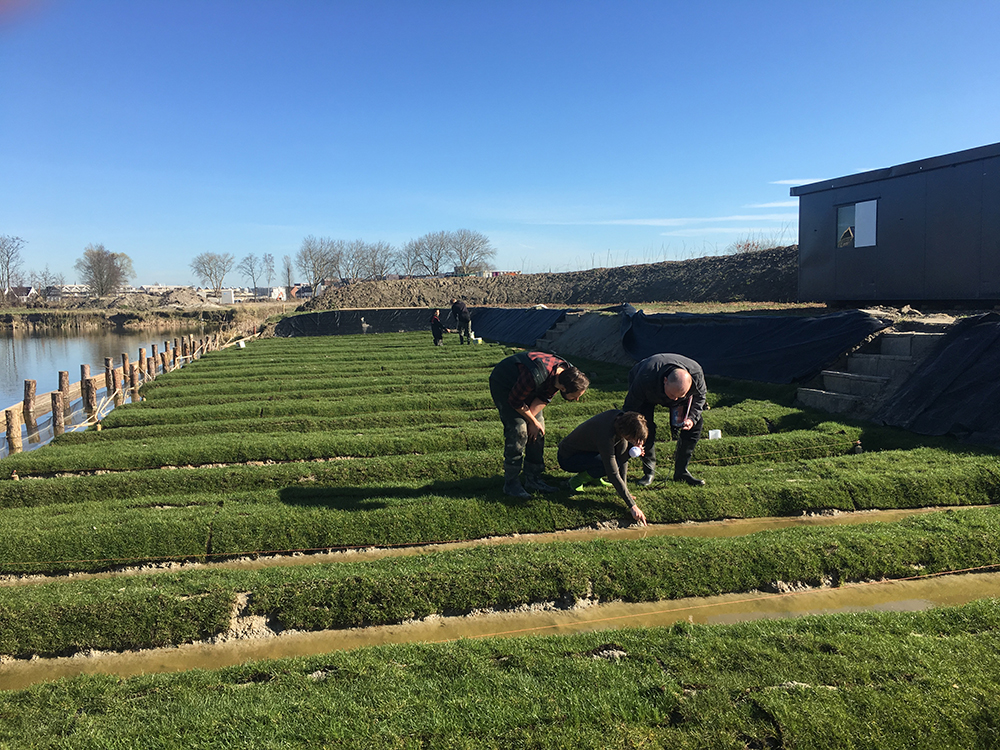

DNA waterscan in the Living Lab
In collaboration with the Institute of Environmental Sciences (CML), Beentjes also tested the eDNA technique in the ditches of the Levend Lab In this outdoor laboratory, biologists study the effects of harmful substances on aquatic life in a natural environment. 'We investigated whether eDNA is suitable for measuring the negative effects of artificial fertiliser and insecticides,' says Beentjes. 'We could indeed clearly see the negative effects on aquatic life. Because we also looked at bacteria and plankton, we also got much more information than we would have had if we had used only the traditional methods.'
How can we use DNA techniques?
Although the techniques are yet to be perfected, Beentjes thinks that they can already be put to good use. 'We can use DNA scans to monitor a lot of places quickly. We can use them as quick scans, to get an indication of the places where more research is needed. We can also pay better attention to animals that we cannot properly visualise using the traditional method, such as plankton and micro-organisms (See box Living Lab). With eDNA we can have a much broader view of the ecosystem!'
